SimPy Model for elective orthopaedic theatre and ward planning#
The aim of this discrete-event simulation (DES) model is to support orthopaedic elective theatre and ward planning.
The DES model enables experimentation with the theatre schedule, number of beds, lengths-of-stay for five orthopaedic elective surgery types, the proportion of patients with a delayed discharge, and the length of delay.
The simulation model is intended to support capacity planning of orthopaedic elective services by identifying a balance of capacity across theatres and beds and predicting the impact of productivity measures on capacity requirements. It is applicable beyond the study site and can be adapted for other specialties.
1. Packages#
The model is provided with a conda virtual environment hep_env
import simpy
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import itertools
import arrow
import random
import math
import warnings
2. Baseline parameters#
2.1 Initialise start date#
Initialise start date to a Monday and define it to be equal to the simulation time This allows monitoring of weekdays
Defined in days: Monday = 0
start = arrow.get('2022-06-27')
# start on a monday
env = simpy.Environment()
2.2 Fitted baseline parameters for NBT based on electronic health records 2016-2019#
Primary: primary hip, primary knee, primary uni-compartmental knee
Revision: revision hip, revision knee
delay_los count 529.000000
*mean 16.521739
*std 15.153132
*min 2.000000
*25% 8.000000
*50% 13.000000
*75% 19.000000
*max 156.000000
primary_knee count 2150.000000
*mean 4.651163
*std 2.828129
*min 0.000000
*25% 3.000000
*50% 4.000000
*75% 6.000000
*max 42.000000
uni_knee count 668.000000
*mean 2.914671
*std 2.136334
*min 0.000000
*25% 2.000000
*50% 3.000000
*75% 4.000000
*max 16.000000
revise_knee count 340.000000
*mean 7.194118
*std 7.598554
*min 0.000000
*25% 3.000000
*50% 5.000000
*75% 9.000000
*max 63.000000
primary_hip count 2820.000000
*mean 4.433333
*std 2.949526
*min 0.000000
*25% 3.000000
*50% 4.000000
*75% 5.000000
*max 44.000000
revise_hip count 406.000000
*mean 6.908867
*std 6.965812
*min 0.000000
*25% 3.000000
*50% 5.000000
*75% 8.000000
*max 86.000000
2.3 Baseline parameters#
# ward parameters
DEFAULT_PRIMARY_HIP_MEAN_LOS = 4.433333
DEFAULT_PRIMARY_KNEE_MEAN_LOS = 4.651163
DEFAULT_REVISION_HIP_MEAN_LOS = 6.908867
DEFAULT_REVISION_KNEE_MEAN_LOS = 7.194118
DEFAULT_UNICOMPART_KNEE_MEAN_LOS = 2.914671
DEFAULT_PRIMARY_HIP_SD_LOS = 2.949526
DEFAULT_PRIMARY_KNEE_SD_LOS = 2.828129
DEFAULT_REVISION_HIP_SD_LOS = 6.965812
DEFAULT_REVISION_KNEE_SD_LOS = 7.598554
DEFAULT_UNICOMPART_KNEE_SD_LOS = 2.136334
DEFAULT_DELAY_POST_LOS_MEAN = 16.521739
DEFAULT_DELAY_POST_LOS_SD = 15.153132
DEFAULT_PROB_WARD_DELAY = 0.076
# ward resources
DEFAULT_NUMBER_BEDS = 40
DEFAULT_PRIMARY_DICT = {1: 'p_hip', 2: 'p_knee', 3: 'uni_knee'}
DEFAULT_REVISION_DICT = {1: 'r_hip', 2: 'r_knee'}
DEFAULT_PRIMARY_PROB = [0.51, 0.38, 0.11]
DEFAULT_REVISION_PROB = [0.55, 0.45]
SET_WEEKDAY = ['Monday', 'Tuesday', 'Wednesday', 'Thursday',
'Friday', 'Saturday', 'Sunday']
SET_SESSIONS_PER_WEEKDAY = {'Monday': 3, 'Tuesday': 3, 'Wednesday': 3, 'Thursday': 3,
'Friday': 3, 'Saturday': 0, 'Sunday': 0}
SET_SESSIONS_PER_WEEKDAY_LIST = list(SET_SESSIONS_PER_WEEKDAY.values())
SET_ALLOCATION = {'Monday': ['2P_or_1R', '2P_or_1R', '1P'],
'Tuesday': ['2P_or_1R', '2P_or_1R', '1P'],
'Wednesday': ['2P_or_1R', '2P_or_1R', '1P'],
'Thursday': ['2P_or_1R', '2P_or_1R', '1P'],
'Friday': ['2P_or_1R', '2P_or_1R', '1P'],
'Saturday': [],
'Sunday': []}
SET_THEATRES_PER_WEEKDAY = {'Monday': 4, 'Tuesday': 4, 'Wednesday': 4, 'Thursday': 4,
'Friday': 4, 'Saturday': 0, 'Sunday': 0}
# simulation parameters
DEFAULT_NUMBER_OF_RUNS = 30
DEFAULT_RESULTS_COLLECTION_PERIOD = 42
results_collection_period = 70
DEFAULT_WARM_UP_PERIOD = 7
default_rng_set = None
# for returning results per day
first_obs = 1
interval = 1
TRACE = False
# extreme schedule generated for testing
new_schedule = pd.read_csv('Test_data/new_schedule.csv')
# empirical los distributions for patients not delayed
los_data = pd.read_csv('Test_data/los_spells_no_zero.csv')
#convert to np arrays for sampling
primary_knee_los_data = los_data['Primary Knee'].dropna().to_numpy()
unicomp_knee_los_data = los_data['Unicompart Knee'].dropna().to_numpy()
revision_knee_los_data = los_data['Revision Knee'].dropna().to_numpy()
primary_hip_los_data = los_data['Primary Hip'].dropna().to_numpy()
revision_hip_los_data = los_data['Revision Hip'].dropna().to_numpy()
#optional use of empirical data to sample los distributions
use_empirical_data = False
# later for translation: ignore this - testing
primary_dict = {1: 'p_hip', 2: 'p_knee', 3: 'uni_knee'}
revision_dict = {1: 'r_hip', 2: 'r_knee'}
primary_prop = np.random.choice(np.arange(1, 4), p=[0.4, 0.4, 0.2])
revision_prop = np.random.choice(np.arange(1, 3), p=[0.6, 0.4])
def vec_tran(prop, dict):
return np.vectorize(dict.__getitem__)(prop)
# trial sample and vectorize by dict key
primary_sample = vec_tran(primary_prop, primary_dict)
3. Trace utility#
def trace(msg):
"""
Used for debugging
If TRUE will return all patient level message output
Default = FALSE
"""
if TRACE:
print(msg)
4. Distribution classes#
Used to control random number sampling
Each distribution has its own stream
Lognormal for lengths-of-stay
Bernoulli for determining lost theatres slots and delayed discharges
class Lognormal:
"""
for creating LoS distributions for each patient type
"""
def __init__(self, mean, stdv, random_seed=None):
self.rng = np.random.default_rng(seed=random_seed)
mu, sigma = self.calc_params(mean, stdv)
self.mu = mu
self.sigma = sigma
def calc_params(self, mean, stdv):
phi = (stdv ** 2 + mean ** 2) ** 0.5
mu = np.log(mean ** 2 / phi)
sigma = (np.log(phi ** 2 / mean ** 2)) ** 0.5
return mu, sigma
def sample(self):
"""
method to generate a sample from the lognormal distribution
"""
return self.rng.lognormal(self.mu, self.sigma)
class Bernoulli:
"""
for pathway branching: slots lost on day,
patients whose LoS is delayed due to downstream processes
"""
def __init__(self, p, random_seed = None):
"""p = prob of drawing a 1"""
self.rng = np.random.default_rng(seed=random_seed)
self.p = p
def sample(self, size = None):
"""
method to generate a sample from the Bernoulli distribution
"""
return self.rng.binomial(n = 1, p = self.p, size = size)
class Gamma:
"""
sensitivity analysis on LoS distributions for each patient type
"""
def __init__(self, mean, stdv, random_seed = None):
self.rng = np.random.default_rng(seed = random_seed)
scale, shape = self.calc_params(mean, stdv)
self.scale = scale
self.shape = shape
def calc_params(self, mean, stdv):
scale = (stdv **2) / mean
shape = (stdv **2) / (scale **2)
return scale, shape
def sample(self, size = None):
"""
method to generate a sample from the gamma distribution
"""
return self.rng.gamma(self.shape, self.scale, size = size)
class Empirical:
"""
for los distributions not fitting statistical distributions
losdata: los per procedure type
"""
def __init__(self, losdata, random_seed = None):
self.rng = np.random.default_rng(seed = random_seed)
self.losdata = losdata
def sample(self, size = None):
"""
method to generate a sample from empirical distribution
"""
return self.rng.choice(self.losdata, size=None, replace=True)
4. Theatre schedule#
Generates a theatre schedule from baseline settings, and optionally for a user-defined schedule.
Baseline settings:
4 theatres (2-6)
5 day/week (5-7)
Each theatre has three sessions per day:
Morning: 1 revision OR 2 primary
Afternoon: 1 revision OR 2 primary
Evening: 1 primary
class Schedule:
"""
Creates theatre schedule according to rules
"""
def __init__(self, weekday=SET_WEEKDAY,
allocation=SET_ALLOCATION,
sessions_per_weekday=SET_SESSIONS_PER_WEEKDAY,
sessions_per_weekday_list=SET_SESSIONS_PER_WEEKDAY_LIST,
theatres_per_weekday=SET_THEATRES_PER_WEEKDAY):
"""
parameters used to create schedule defined in 'scenarios class'
"""
self.weekday=weekday
self.allocation=allocation
self.sessions_per_weekday=sessions_per_weekday
self.sessions_per_weekday_list=sessions_per_weekday_list
self.theatres_per_weekday=theatres_per_weekday
def create_schedule(self,weekday, sessions_per_weekday_list, allocation, theatres_per_weekday):
"""
Arguments needed:
*weekday: a list of weekdays
*sessions_per_weekday: a list of integers representing the number of sessions per weekday
*allocation: a dictionary where the keys are the weekdays and the values are lists of
allocations for each session
*theatres_per_weekday: a dictionary where the keys are the weekdays and the values are
integers representing the number of theatres per weekday
Returns a dictionary where the keys are the weekdays and the values are lists
of lists of allocations for each theatre for each session.
"""
schedule = {}
for day, num_sessions in zip(weekday, sessions_per_weekday_list):
schedule[day] = []
for theatre in range(theatres_per_weekday[day]):
schedule[day].append([])
for session in range(num_sessions):
if allocation[day][session] == '1P':
schedule[day][theatre].append({'primary': 1})
elif allocation[day][session] == '1R':
schedule[day][theatre].append({'revision': 1})
elif allocation[day][session] == '2P':
schedule[day][theatre].append({'primary': 2})
elif allocation[day][session] == '2P_or_1R':
if random.random() > 0.5:
schedule[day][theatre].append({'primary': 2})
else:
schedule[day][theatre].append({'revision': 1})
return schedule
def daily_counts(self,day_data):
"""
day_data: called in week_schedule() function, day_data is a sample weekly dictionary from create_schedule()
Convert dict to a pandas DataFrame with 'primary' and 'revision' as columns
and days of the week as the index, populated with the total count of 'primary' and 'revision' in each day.
Returns a one week schedule
"""
#day_data = create_schedule(weekday, sessions_per_weekday, allocation, theatres_per_weekday)
primary_slots = 0
revision_slots = 0
for value in day_data:
if value:
for sub_value in value:
if 'primary' in sub_value:
primary_slots += sub_value['primary']
if 'revision' in sub_value:
revision_slots += sub_value['revision']
return [primary_slots, revision_slots]
def week_schedule(self):
"""
samples a weekly dictionary of theatres, sessions, and surgeries from create_schedule()
counts daily number or primary and revision surgeries needed using daily_counts()
and converts to a dataframe
"""
week_sched = pd.DataFrame(columns=['Primary_slots', 'Revision_slots'])
day_data = self.create_schedule(self.weekday, self.sessions_per_weekday_list,
self.allocation, self.theatres_per_weekday)
for key, value in day_data.items():
week_sched.loc[key] = self.daily_counts(value)
week_sched = week_sched.reset_index()
week_sched.rename(columns = {'index':'Day'}, inplace = True)
return week_sched
def theatre_capacity(self):
length_sched = int(round(2*(DEFAULT_WARM_UP_PERIOD+DEFAULT_RESULTS_COLLECTION_PERIOD)/7, 0))
DEFAULT_SCHEDULE_AVAIL = pd.DataFrame()
for week in range(length_sched):
single_random_week = self.week_schedule()
DEFAULT_SCHEDULE_AVAIL = pd.concat([DEFAULT_SCHEDULE_AVAIL, single_random_week],axis=0)
return DEFAULT_SCHEDULE_AVAIL.reset_index()
# class Save:
# schedule = Schedule()
# def __init__(self, schedule):
# self.schedule = schedule
# schedule_avail3 = schedule.theatre_capacity()
# print(schedule_avail3.head(7))
# schedule_avail3.to_csv('data/new_schedule_checking.csv')
5. Model parameterisation#
A scenarios class containing all parameters that can be varied in the model.
Used for setting up scenarios, the scenarios class contains baseline parameters which can be changed at runtime.
class Scenario:
"""
Holds LoS dists for each patient type
Holds delay dists
Holds prob of delay, prob of same day dist
Holds resources: beds
Passed to hospital model and process classes
"""
def __init__(self, schedule, schedule_avail=None,
random_number_set=default_rng_set,
primary_hip_mean_los=DEFAULT_PRIMARY_HIP_MEAN_LOS,
primary_knee_mean_los=DEFAULT_PRIMARY_KNEE_MEAN_LOS,
revision_hip_mean_los=DEFAULT_REVISION_HIP_MEAN_LOS,
revision_knee_mean_los=DEFAULT_REVISION_KNEE_MEAN_LOS,
unicompart_knee_mean_los=DEFAULT_UNICOMPART_KNEE_MEAN_LOS,
delay_post_los_mean=DEFAULT_DELAY_POST_LOS_MEAN,
prob_ward_delay=DEFAULT_PROB_WARD_DELAY,
n_beds=DEFAULT_NUMBER_BEDS,
primary_hip_sd_los=DEFAULT_PRIMARY_HIP_SD_LOS,
primary_knee_sd_los=DEFAULT_PRIMARY_KNEE_SD_LOS,
revision_hip_sd_los=DEFAULT_REVISION_HIP_SD_LOS,
revision_knee_sd_los=DEFAULT_REVISION_KNEE_SD_LOS,
unicompart_knee_sd_los=DEFAULT_UNICOMPART_KNEE_SD_LOS,
delay_post_los_sd=DEFAULT_DELAY_POST_LOS_SD,
primary_dict=DEFAULT_PRIMARY_DICT,
revision_dict=DEFAULT_REVISION_DICT,
primary_prob=DEFAULT_PRIMARY_PROB,
revision_prob=DEFAULT_REVISION_PROB,
primary_knee_los_data = primary_knee_los_data,
unicompart_knee_los_data = unicomp_knee_los_data,
revision_knee_los_data = revision_knee_los_data,
primary_hip_los_data = primary_hip_los_data,
revision_hip_los_data = revision_hip_los_data):
"""
controls initial seeds of each RNS used in model
"""
self.schedule = schedule
if schedule_avail is None:
self.schedule_avail = schedule.theatre_capacity()
else:
self.schedule_avail = schedule_avail
#self.schedule_avail = schedule_avail
self.random_number_set = random_number_set
self.primary_hip_mean_los = primary_hip_mean_los
self.primary_knee_mean_los = primary_knee_mean_los
self.revision_hip_mean_los = revision_hip_mean_los
self.revision_knee_mean_los = revision_knee_mean_los
self.unicompart_knee_mean_los = unicompart_knee_mean_los
self.n_beds = n_beds
self.prob_ward_delay = prob_ward_delay
self.primary_hip_sd_los = primary_hip_sd_los
self.primary_knee_sd_los = primary_knee_sd_los
self.revision_hip_sd_los = revision_hip_sd_los
self.revision_knee_sd_los = revision_knee_sd_los
self.unicompart_knee_sd_los = unicompart_knee_sd_los
self.delay_post_los_mean = delay_post_los_mean
self.delay_post_los_sd = delay_post_los_sd
self.primary_dict = primary_dict
self.revision_dict = revision_dict
self.primary_prob = primary_prob
self.revision_prob = revision_prob
self.primary_knee_los_data = primary_knee_los_data
self.unicompart_knee_los_data = unicompart_knee_los_data
self.revision_knee_los_data = revision_knee_los_data
self.primary_hip_los_data = primary_hip_los_data
self.revision_hip_los_data = revision_hip_los_data
self.init_sampling()
def set_random_no_set(self, random_number_set):
"""
controls random sampling for each distribution used in simulations"""
self.random_number_set = random_number_set
self.init_sampling()
def init_sampling(self):
"""
distribs used in model and initialise seed"""
rng_streams = np.random.default_rng(self.random_number_set)
self.seeds = rng_streams.integers(0,99999999999, size = 20)
####### Distributions ########
# LNorm LoS distribution for each surgery patient type
self.primary_hip_dist = Lognormal(self.primary_hip_mean_los, self.primary_hip_sd_los,
random_seed=self.seeds[0])
self.primary_knee_dist = Lognormal(self.primary_knee_mean_los, self.primary_knee_sd_los,
random_seed=self.seeds[1])
self.revision_hip_dist = Lognormal(self.revision_hip_mean_los, self.revision_hip_sd_los,
random_seed=self.seeds[2])
self.revision_knee_dist = Lognormal(self.revision_knee_mean_los, self.revision_knee_sd_los,
random_seed=self.seeds[3])
self.unicompart_knee_dist = Lognormal(self.unicompart_knee_mean_los, self.unicompart_knee_sd_los,
random_seed=self.seeds[4])
# Empirical LoS distributions for each surgery patient type
self.primary_hip_dist_emp = Empirical(self.primary_hip_los_data,
random_seed=self.seeds[5])
self.primary_knee_dist_emp = Empirical(self.primary_knee_los_data,
random_seed=self.seeds[6])
self.revision_hip_dist_emp = Empirical(self.revision_hip_los_data,
random_seed=self.seeds[7])
self.revision_knee_dist_emp = Empirical(self.revision_knee_los_data,
random_seed=self.seeds[8])
self.unicompart_knee_dist_emp = Empirical(self.unicompart_knee_los_data,
random_seed=self.seeds[9])
# distribution for delayed LoS
self.los_delay_dist = Lognormal(self.delay_post_los_mean, self.delay_post_los_sd,
random_seed=self.seeds[10])
#probability of having LoS delayed on ward
self.los_delay = Bernoulli(self.prob_ward_delay, random_seed=self.seeds[11])
def number_slots(self, schedule_avail):
"""
convert to np arrays for each surgery type for patient generators
"""
self.schedule_avail_primary = self.schedule_avail['Primary_slots'].to_numpy()
self.schedule_avail_revision = self.schedule_avail['Revision_slots'].to_numpy()
return(self.schedule_avail_primary, self.schedule_avail_revision)
def primary_types(self,prob):
"""
randomly select primary surgical type from custom distribution: primary_prop
prob = primary_prop
used for generating primary patients of each surgical type
"""
self.primary_surgery = np.random.choice(np.arange(1,4), p=prob)
return(self.primary_surgery)
def revision_types(self,prob):
"""
randomly select revision surgical type from custom distribution: revision_prop
prob = revision_prop
used for generating revision patients of each surgical type
"""
self.revision_surgery = np.random.choice(np.arange(1,3), p=prob)
return(self.revision_surgery)
def label_types(self, prop, dict):
"""
return label for each surgery type
"""
return np.vectorize(dict.__getitem__)(prop)
6. Patient pathways process logic#
Patient journeys for two classes: PrimaryPatient and RevisionPatient
class PrimaryPatient:
"""
The process a patient needing primary hip or knee surgery will undergo
from scheduled admission for surgery to discharge
day = simulation day
id = patient id
args: Scenario parameter class
"""
def __init__(self, day, id, env, args):
self.day = day
self.id = id
self.env = env
self.args = args
self.arrival = -np.inf
self.queue_beds = -np.inf
self.primary_los = 0
self.total_time = -np.inf
self.depart = -np.inf
self.lost_slots_bool = False
self.delayed_los_bool = False
self.weekday = 0
self.patient_class = 'primary'
def service(self):
"""
Arrive according to theatres schedule
Some patients will leave on day of surgery and the slot is lost
Some patients will have their surgery cancelled due to lack of beds
Otherwise, patient is admitted and stays in a bed
Some patients will have a post-bed request delay to their LoS
Patient is discharged
"""
self.arrival = self.env.now
self.patient_class = 'primary'
self.weekday = start.shift(days=self.env.now).weekday()
# set los for primary surgery types
self.types = int(self.args.primary_types(self.args.primary_prob))
if self.types == 1:
if use_empirical_data:
self.primary_los = self.args.primary_hip_dist_emp.sample()
else:
self.primary_los = self.args.primary_hip_dist.sample()
self.primary_label = 'p_hip'
elif self.types == 2:
if use_empirical_data:
self.primary_los = self.args.primary_knee_dist_emp.sample()
else:
self.primary_los = self.args.primary_knee_dist.sample()
self.primary_label = 'p_knee'
else:
if use_empirical_data:
self.primary_los = self.args.unicompart_knee_dist_emp.sample()
else:
self.primary_los = self.args.unicompart_knee_dist.sample()
self.primary_label = 'uni_knee'
#vectorize according to dict key to get surgical type
#self.primary_label = self.args.label_types(primary_prop, primary_dict)
#sample if need for delayed discharge
self.need_for_los_delay = self.args.los_delay.sample()
#Patients who have a delayed discharge follow this pathway
if self.need_for_los_delay:
#request a bed on ward - if none available within 0.5-1 day, patient has surgery cancelled
with self.args.beds.request() as req:
admission = random.uniform(0.5,1)
admit = yield req | self.env.timeout(admission)
if req in admit:
"""record queue time for primary patients -- if > admission,
this patient will leave the system and the slot is lost"""
self.queue_beds = self.env.now - self.arrival
trace(f'primary patient {self.id} {self.primary_label}'
f'has been allocated a bed at {self.env.now:.3f}'
f'and queued for {self.queue_beds:.3f}')
self.primary_los = args.los_delay_dist.sample()
yield self.env.timeout(self.primary_los)
self.lost_slots_bool = False
self.delayed_los_bool = True
self.depart = self.env.now
trace(f'los of primary patient {self.id} completed at {self.env.now:.3f}')
self.total_time = self.env.now - self.arrival
trace(f'primary patient {self.id} {self.primary_label}'
f'total los = {self.total_time:.3f} with delayed discharge')
else:
#patient had to leave as no beds were available on ward
self.no_bed_cancellation = self.env.now - self.arrival
trace(f'primary patient {self.id} {self.primary_label}'
f'had surgery cancelled after {self.no_bed_cancellation:.3f}')
self.queue_beds = self.env.now - self.arrival
self.total_time = self.env.now - self.arrival
self.primary_los = 0
self.lost_slots_bool = True
self.delayed_los_bool = False
self.depart = self.env.now
trace(f'primary patient {self.id} {self.primary_label}'
f'recorded {self.lost_slots_bool}')
#no delayed los
else:
#request a bed on ward - if none available within 0.5-1 day, patient has surgery cancelled
with self.args.beds.request() as req:
admission = random.uniform(0.5,1)
admit = yield req | self.env.timeout(admission)
self.no_bed_cancellation = self.env.now - self.arrival
if req in admit:
#record queue time for primary patients -- if >1, this patient will leave the system and the slot is lost
self.queue_beds = self.env.now - self.arrival
trace(f'primary patient {self.id} {self.primary_label}'
f'has been allocated a bed at {self.env.now:.3f}'
f'and queued for {self.queue_beds:.3f}')
self.primary_los = self.primary_los
yield self.env.timeout(self.primary_los)
self.lost_slots_bool = False
self.delayed_los_bool = False
self.depart = self.env.now
trace(f'los of primary patient {self.id} {self.primary_label}'
f'completed at {self.env.now:.3f}')
self.total_time = self.env.now - self.arrival
trace(f'primary patient {self.id} {self.primary_label}'
f'total los = {self.total_time:.3f}')
else:
#patient had to leave as no beds were available on ward
trace(f'primary patient {self.id} {self.primary_label}'
f'had surgery cancelled after {self.no_bed_cancellation:.3f}')
self.queue_beds = self.env.now - self.arrival
self.total_time = self.env.now - self.arrival
self.primary_los = 0
self.lost_slots_bool = True
self.delayed_los_bool = False
self.depart = self.env.now
trace(f'primary patient {self.id} {self.primary_label}'
f'recorded {self.lost_slots_bool}')
class RevisionPatient:
"""
The process a patient needing revision hip or knee surgery will undergo
from scheduled admission for surgery to discharge
day = simulation day
id = patient id
args: Scenario parameter class
"""
def __init__(self, day, id, env, args):
self.day = day
self.id = id
self.env = env
self.args = args
self.arrival = -np.inf
self.queue_beds = -np.inf
self.revision_los = 0
self.total_time = -np.inf
self.depart = -np.inf
self.lost_slots_bool = False
self.delayed_los_bool = False
self.weekday = 0
self.patient_class = 'revision'
def service(self):
"""
Arrive according to theatres schedule
Some patients will leave on day of surgery and the slot is lost
Some patients will have their surgery cancelled due to lack of beds
Otherwise, patient is admitted and stays in a bed
Some patients will have a post-bed request delay to their LoS
Patient is discharged
"""
self.arrival = self.env.now
self.patient_class = 'revision'
self.weekday = start.shift(days=self.env.now).weekday()
# set los for revision surgery types
self.types = int(self.args.revision_types(self.args.revision_prob))
if self.types == 1:
if use_empirical_data:
self.revision_los = self.args.revision_hip_dist_emp.sample()
else:
self.revision_los = self.args.revision_hip_dist.sample()
self.revision_label = 'r_hip'
else:
if use_empirical_data:
self.revision_los = self.args.revision_knee_dist_emp.sample()
else:
self.revision_los = self.args.revision_knee_dist.sample()
self.revision_label = 'r_knee'
#vectorize according to dict key to get surgical type
#self.revision_label = self.args.label_types(revision_prop, revision_dict)
#sample if need for delayed discharge
self.need_for_los_delay = self.args.los_delay.sample()
if self.need_for_los_delay:
#request bed on ward - if none available within 0.5-1 day, patient has surgery cancelled
with self.args.beds.request() as req:
admission = random.uniform(0.5, 1)
admit = yield req | self.env.timeout(admission)
if req in admit:
#record queue time for primary patients -- if >admission, this patient will leave the system and the slot is lost
self.queue_beds = self.env.now - self.arrival
trace(f'revision patient {self.id} {self.revision_label}'
f'has been allocated a bed at {self.env.now:.3f}'
f'and queued for {self.queue_beds:.3f}')
self.revision_los = args.los_delay_dist.sample()
yield self.env.timeout(self.revision_los)
self.lost_slots_bool = False
self.delayed_los_bool = True
self.depart = self.env.now
trace(f'los of revision patient {self.id} {self.revision_label}'
f'completed at {self.env.now:.3f}')
self.total_time = self.env.now - self.arrival
trace(f'revision patient {self.id} {self.revision_label}'
f'total los = {self.total_time:.3f} with delayed discharge')
else:
#patient had to leave as no beds were available on ward
self.no_bed_cancellation = self.env.now - self.arrival
trace(f'revision patient {self.id}'
f'had surgery cancelled after {self.no_bed_cancellation:.3f}')
self.queue_beds = self.env.now - self.arrival
self.total_time = self.env.now - self.arrival
self.revision_los = 0
self.lost_slots_bool = True
self.delayed_los_bool = False
self.depart = self.env.now
trace(f'revision patient {self.id} {self.revision_label}'
f'recorded {self.lost_slots_bool}')
#no need for delayed discharge
else:
#request bed on ward - if none available within 0.5-1 day, patient has surgery cancelled
with self.args.beds.request() as req:
admission = random.uniform(0.5, 1)
admit = yield req | self.env.timeout(admission)
self.no_bed_cancellation = self.env.now - self.arrival
if req in admit:
#record queue time for primary patients -- if >1, this patient will leave the system and the slot is lost
self.queue_beds = self.env.now - self.arrival
trace(f'revision patient {self.id} {self.revision_label}'
f'has been allocated a bed at {self.env.now:.3f}'
f'and queued for {self.queue_beds:.3f}')
self.revision_los = self.revision_los
yield self.env.timeout(self.revision_los)
self.lost_slots_bool = False
self.delayed_los_bool = False
self.depart = self.env.now
trace(f'los of revision patient {self.id} completed at {self.env.now:.3f}')
self.total_time = self.env.now - self.arrival
trace(f'revision patient {self.id} total los = {self.total_time:.3f}')
else:
#patient had to leave as no beds were available on ward
trace(f'revision patient {self.id} {self.revision_label}'
f'had surgery cancelled after {self.no_bed_cancellation:.3f}')
self.queue_beds = self.env.now - self.arrival
self.total_time = self.env.now - self.arrival
self.revision_los = 0
self.lost_slots_bool = True
self.delayed_los_bool = False
self.depart = self.env.now
trace(f'revision patient {self.id} {self.revision_label}'
f'recorded {self.lost_slots_bool}')
7. The model class#
The Hospital class generates primary and revision patients and implements a method to run the model.
Collects patient-level results and audits daily results
class Hospital:
"""
The orthopaedic hospital model
"""
def __init__(self, args):
self.env = simpy.Environment()
self.args = args
self.init_resources()
#patient generator lists
self.patients = []
self.primary_patients = []
self.revision_patients = []
self.primary_patients_id = []
self.revision_patients_id = []
self.cum_primary_patients = []
self.cum_revision_patients = []
self.DEFAULT_RESULTS_COLLECTION_PERIOD = None
self.summary_results = None
self.audit_interval = interval
#lists used for daily audit_frame for summary results per day
self.audit_time = []
self.audit_day_of_week = []
self.audit_beds_used_primary = []
self.audit_beds_used_revision = []
self.audit_beds_used = []
self.audit_primary_arrival = []
self.audit_revision_arrival = []
self.audit_primary_queue_beds = []
self.audit_revision_queue_beds = []
self.audit_primary_los = []
self.audit_revision_los = []
self.results = pd.DataFrame()
def audit_frame(self):
"""
Dataframe with results summarised per day
"""
self.results = pd.DataFrame({'sim_time':self.audit_time,
'weekday': self.audit_day_of_week,
'bed_utilisation_primary': self.audit_beds_used_primary,
'bed_utilisation_revision': self.audit_beds_used_revision,
'bed_utilisation':self.audit_beds_used,
'primary_arrivals': self.audit_primary_arrival,
'revision_arrivals': self.audit_revision_arrival,
'primary_bed_queue': self.audit_primary_queue_beds,
'revision_bed_queue': self.audit_revision_queue_beds,
'primary_mean_los': self.audit_primary_los,
'revision_mean_los': self.audit_revision_los
})
def patient_results(self):
"""
Dataframes to hold individual results per patient per day per run
Attributes from patient classes
"""
results_primary_pt = pd.DataFrame({'Day':np.array([getattr(p, 'day') for p in self.cum_primary_patients]),
'weekday':np.array([getattr(p, 'weekday') for p in self.cum_primary_patients]),
'ID':np.array([getattr(p, 'id') for p in self.cum_primary_patients]),
'arrival time':np.array([getattr(p, 'arrival') for p in self.cum_primary_patients]),
'patient class':np.array([getattr(p, 'patient_class') for p in self.cum_primary_patients]),
'surgery type':np.array([getattr(p, 'primary_label') for p in self.cum_primary_patients]),
'lost slots':np.array([getattr(p, 'lost_slots_bool') for p in self.cum_primary_patients]),
'queue time':np.array([getattr(p, 'queue_beds') for p in self.cum_primary_patients]),
'los':np.array([getattr(p, 'primary_los') for p in self.cum_primary_patients]),
'delayed discharge':np.array([getattr(p, 'delayed_los_bool') for p in self.cum_primary_patients]),
'depart':np.array([getattr(p, 'depart') for p in self.cum_primary_patients])
})
results_revision_pt = pd.DataFrame({'Day':np.array([getattr(p, 'day') for p in self.cum_revision_patients]),
'ID':np.array([getattr(p, 'id') for p in self.cum_revision_patients]),
'weekday':np.array([getattr(p, 'weekday') for p in self.cum_revision_patients]),
'arrival time':np.array([getattr(p, 'arrival') for p in self.cum_revision_patients]),
'patient class':np.array([getattr(p, 'patient_class') for p in self.cum_revision_patients]),
'surgery type':np.array([getattr(p, 'revision_label') for p in self.cum_revision_patients]),
'lost slots':np.array([getattr(p, 'lost_slots_bool') for p in self.cum_revision_patients]),
'queue time':np.array([getattr(p, 'queue_beds') for p in self.cum_revision_patients]),
'los':np.array([getattr(p, 'revision_los') for p in self.cum_revision_patients]),
'delayed discharge':np.array([getattr(p, 'delayed_los_bool') for p in self.cum_revision_patients]),
'depart':np.array([getattr(p, 'depart') for p in self.cum_revision_patients])
})
return(results_primary_pt, results_revision_pt)
def plots(self):
"""
plot results at end of run
"""
def perform_audit(self):
"""
Results per day
monitor ED each day and return daily results for metrics in audit_frame
"""
yield self.env.timeout(DEFAULT_WARM_UP_PERIOD)
while True:
#simulation time
t = self.env.now
self.audit_time.append(t)
#weekday
self.audit_day_of_week.append(start.shift(days=self.env.now -1).weekday())
########## bed utilisation - primary, revision, total
primary_beds = self.args.beds.count in self.cum_primary_patients
(self.audit_beds_used_primary.append(primary_beds / self.args.n_beds))
revision_beds = self.args.beds.count in self.cum_revision_patients
(self.audit_beds_used_revision.append(revision_beds / self.args.n_beds))
(self.audit_beds_used.append(self.args.beds.count / self.args.n_beds))
########### lost slots
patients = self.cum_revision_patients + self.cum_primary_patients
# deal with lost slots on zero arrival days
"""
lost_slots = []
def zero_days(ls):
if not zero_days:
return 1
else:
return 0
ls = (np.array([getattr(p, 'lost_slots_int') for p in patients]))
if zero_days(ls):
lost_slots = 0
else:
lost_slots = len(np.array([getattr(p,'lost_slots_int') for p in patients / len(patients)
self.audit_slots_lost.append(lost_slots)
"""
######### arrivals
pp = len(np.array([p.id for p in self.cum_primary_patients]))
rp = len(np.array([p.id for p in self.cum_revision_patients]))
self.audit_primary_arrival.append(len(self.primary_patients))
self.audit_revision_arrival.append(len(self.revision_patients))
#queue times
primary_q = np.array([getattr(p, 'queue_beds') for p in self.cum_primary_patients
if getattr(p, 'queue_beds') > -np.inf]).mean()
self.audit_primary_queue_beds.append(primary_q)
revision_q = np.array([getattr(p, 'queue_beds') for p in self.cum_revision_patients
if getattr(p, 'queue_beds') > -np.inf]).mean()
self.audit_revision_queue_beds.append(revision_q)
#mean lengths of stay
primarylos = np.array([getattr(p, 'primary_los') for p in self.cum_primary_patients
if getattr(p, 'primary_los') > -np.inf]).mean().round(2)
self.audit_primary_los.append(primarylos)
revisionlos = np.array([getattr(p, 'revision_los') for p in self.cum_revision_patients
if getattr(p, 'revision_los') > -np.inf]).mean().round(2)
self.audit_revision_los.append(revisionlos)
yield self.env.timeout(self.audit_interval)
def init_resources(self):
"""
ward beds initialised and stored in args
"""
self.args.beds = simpy.Resource(self.env,
capacity=self.args.n_beds)
def run(self, results_collection = DEFAULT_RESULTS_COLLECTION_PERIOD+DEFAULT_WARM_UP_PERIOD):
"""
single run of model
"""
self.env.process(self.patient_arrivals_generator_primary())
self.env.process(self.patient_arrivals_generator_revision())
self.env.process(self.perform_audit())
self.results_collection = results_collection
self.env.run(until=results_collection)
audit_frame = self.audit_frame()
return audit_frame
def patient_arrivals_generator_primary(self):
"""
Primary patients arrive according to daily theatre schedule
------------------
"""
#sched = args.number_slots(self.args.schedule_avail)[0]
sched = self.args.schedule_avail['Primary_slots']
pt_count = 1
for day in range(len(sched)):
primary_arrivals = sched[day]
trace(f'--------- {primary_arrivals} primary patients are scheduled on Day {day} -------')
for i in range(primary_arrivals):
new_primary_patient = PrimaryPatient(day, pt_count, self.env, self.args)
self.cum_primary_patients.append(new_primary_patient)
self.primary_patients.append(new_primary_patient)
#for debuggng
self.primary_patients_id.append(new_primary_patient.id)
trace(f'primary patient {pt_count} arrived on day {day:.3f}')
self.env.process(new_primary_patient.service())
pt_count += 1
trace(f'primary ids: {self.primary_patients_id}')
yield self.env.timeout(1)
self.primary_patients *= 0
def patient_arrivals_generator_revision(self):
"""
Revision patients arrive according to daily theatre schedule
------------------
"""
sched = args.number_slots(self.args.schedule_avail)[1]
sched = self.args.schedule_avail['Revision_slots']
pt_count = 1
for day in range(len(sched)):
revision_arrivals = sched[day]
trace(f'--------- {revision_arrivals} revision patients are scheduled on Day {day} -------')
for i in range(revision_arrivals):
new_revision_patient = RevisionPatient(day, pt_count, self.env, self.args)
self.cum_revision_patients.append(new_revision_patient)
self.revision_patients.append(new_revision_patient)
#for debugging
self.revision_patients_id.append(new_revision_patient.id)
trace(f'revision patient {pt_count} arrived on day {day:.3f}')
self.env.process(new_revision_patient.service())
pt_count += 1
trace(f'revision ids: {self.revision_patients_id}')
yield self.env.timeout(1)
self.revision_patients *= 0
8. Summary results across days and runs#
Overall summary of results across all runs, and across model run time.
Used for validation
class Summary:
"""
summary results across run
"""
def __init__(self, model):
""" model: Hospital """
self.model = model
self.args = model.args
self.summary_results = None
def process_run_results(self):
self.summary_results = {}
#all patients arrived during results collection period
patients = len([p for p in self.model.cum_primary_patients if p.day > DEFAULT_WARM_UP_PERIOD])+\
len([p for p in self.model.cum_revision_patients if p.day > DEFAULT_WARM_UP_PERIOD])
primary_arrivals = len([p for p in self.model.cum_primary_patients if p.day > DEFAULT_WARM_UP_PERIOD])
revision_arrivals = len([p for p in self.model.cum_revision_patients if p.day > DEFAULT_WARM_UP_PERIOD])
#throughput during results collection period
primary_throughput = len([p for p in self.model.cum_primary_patients if (p.total_time > -np.inf)
& (p.day > DEFAULT_WARM_UP_PERIOD)])
revision_throughput = len([p for p in self.model.cum_revision_patients if (p.total_time > -np.inf)
& (p.day > DEFAULT_WARM_UP_PERIOD)])
#mean queues - this also includes patients who renege and therefore have 0 queue
mean_primary_queue_beds = np.array([getattr(p, 'queue_beds') for p in self.model.cum_primary_patients
if getattr(p, 'queue_beds') > -np.inf]).mean()
mean_revision_queue_beds = np.array([getattr(p, 'queue_beds') for p in self.model.cum_revision_patients
if getattr(p, 'queue_beds') > -np.inf]).mean()
#check mean los
mean_primary_los = np.array([getattr(p, 'primary_los') for p in self.model.cum_primary_patients
if getattr(p, 'primary_los') > 0]).mean()
mean_revision_los = np.array([getattr(p, 'revision_los') for p in self.model.cum_revision_patients
if getattr(p, 'revision_los') > 0]).mean()
#bed utilisation primary and revision patients during results collection period
los_primary = np.array([getattr(p,'primary_los') for p in self.model.cum_primary_patients
if (getattr(p, 'primary_los') > -np.inf) & (getattr(p, 'day') > DEFAULT_WARM_UP_PERIOD)]).sum()
mean_primary_bed_utilisation = los_primary / (DEFAULT_RESULTS_COLLECTION_PERIOD * self.args.n_beds)
los_revision = np.array([getattr(p,'revision_los') for p in self.model.cum_revision_patients
if (getattr(p, 'revision_los') > -np.inf) & (getattr(p, 'day') > DEFAULT_WARM_UP_PERIOD)]).sum()
mean_revision_bed_utilisation = los_revision / (DEFAULT_RESULTS_COLLECTION_PERIOD * self.args.n_beds)
self.summary_results = {'arrivals':patients,
'primary_arrivals':primary_arrivals,
'revision_arrivals':revision_arrivals,
'primary_throughput':primary_throughput,
'revision_throughput':revision_throughput,
'primary_queue':mean_primary_queue_beds,
'revision_queue':mean_revision_queue_beds,
'mean_primary_los':mean_primary_los,
'mean_revision_los':mean_revision_los,
'primary_bed_utilisation':mean_primary_bed_utilisation,
'revision_bed_utilisation':mean_revision_bed_utilisation}
def summary_frame(self):
if self.summary_results is None:
self.process_run_results()
df = pd.DataFrame({'1':self.summary_results})
df = df.T
df.index.name = 'rep'
return df
9. Model execution#
A single_run returns three data sets: summary, daily, patient-level
The function multiple_reps calls single_run for the number of replications.
def single_run(scenario, results_collection=DEFAULT_RESULTS_COLLECTION_PERIOD+DEFAULT_WARM_UP_PERIOD, random_no_set=default_rng_set):
"""
summary results for a single run which can be called for multiple runs
1. summary of single run
2. daily audit of mean results per day
3a. primary patient results for one run and all days
3b. revision patient results for one run and all days
"""
scenario.set_random_no_set(random_no_set)
schedule = Schedule()
model = Hospital(scenario)
model.run(results_collection = results_collection)
summary = Summary(model)
#summary results for a single run
#(warmup excluded apart from bed utilisation AND throughput)
summary_df = summary.summary_frame()
#summary per day results for a single run (warmup excluded)
results_per_day = model.results
#patient-level results (includes warmup results)
patient_results = model.patient_results()
return(summary_df, results_per_day, patient_results)
def multiple_reps(scenario, results_collection=DEFAULT_RESULTS_COLLECTION_PERIOD+DEFAULT_WARM_UP_PERIOD,
n_reps=DEFAULT_NUMBER_OF_RUNS):
"""
create dataframes of summary results across multiple runs:
1. summary table per run
2. summary table per run and per day
3a. primary patient results for all days and all runs
3b. revision patient results for all days and all runs
"""
#summary per run for multiple reps
#(warm-up excluded apart from bed utilisation AND throughput)
results = [single_run(scenario, results_collection, random_no_set=rep)[0]
for rep in range(n_reps)]
df_results = pd.concat(results)
df_results.index = np.arange(1, len(df_results)+1)
df_results.index.name = 'rep'
#summary per day per run for multiple reps (warmup excluded)
day_results = [single_run(scenario, results_collection, random_no_set=rep)[1]
for rep in range(n_reps)]
length_run = [*range(1, results_collection-DEFAULT_WARM_UP_PERIOD+1)]
length_reps = [*range(1, n_reps+1)]
run = [rep for rep in length_reps for i in length_run]
df_day_results = pd.concat(day_results)
df_day_results['run'] = run
#patient results for all days and all runs (warmup included)
primary_pt_results = [single_run(scenario, results_collection, random_no_set=rep)[2][0]
for rep in range(n_reps)]
primary_pt_results = pd.concat(primary_pt_results)
revision_pt_results = [single_run(scenario, results_collection, random_no_set=rep)[2][1]
for rep in range(n_reps)]
revision_pt_results = pd.concat(revision_pt_results)
return (df_results, df_day_results, primary_pt_results, revision_pt_results)
9.1 A single run#
Run the model once and check summary results
# a single run
schedule = Schedule()
args = Scenario(schedule)
s_results = single_run(args, random_no_set = 42)
print(repr(s_results[0].T))
rep 1
arrivals 454.000000
mean_primary_los 4.996316
mean_revision_los 6.143514
primary_arrivals 328.000000
primary_bed_utilisation 0.767368
primary_queue 0.316710
primary_throughput 304.000000
revision_arrivals 126.000000
revision_bed_utilisation 0.166362
revision_queue 0.537871
revision_throughput 122.000000
9.2 Multiple independent replications#
Multiple runs of model and unpack results into summary, daily, and patient level (for primary and revision)
args = Scenario(schedule)
results = multiple_reps(args, n_reps=DEFAULT_NUMBER_OF_RUNS)
m_results = results[0]
m_day_results = results[1]
m_primary_pt_results = results[2]
r_revision_pt_results = results[3]
9.3 Plot summary results for multiple runs#
def summary_over_runs(m_results):
"""
summary results for multiple runs
throughput and bed utilisation excludes results warm-up - arrivals include warmup
visualise replications for throughput and utilisation
"""
summ = m_results.mean().round(2)
fig, ax = plt.subplots(4,1, figsize=(12,10))
ax[0].hist(m_results['primary_bed_utilisation']);
ax[0].set_ylabel('Primary bed utilisation')
ax[1].hist(m_results['revision_bed_utilisation']);
ax[1].set_ylabel('Revision bed utilisation')
ax[2].hist(m_results['primary_throughput']);
ax[2].set_ylabel('Primary throughput')
ax[3].hist(m_results['revision_throughput']);
ax[3].set_ylabel('Revision throughput')
return(summ, fig)
#summary_over_runs(m_results)
10. Summary results per day for multiple runs for bed utilisation#
Group by simulation time (day) across all runs
def daily_summ_bed_utilisation(m_day_results):
"""
summarise per day across runs and save to csv in case of further analysis
print bed utilisation plot
warm-up results are excluded at runtime
"""
m_day_results_ts = m_day_results.groupby(['sim_time']).mean()
m_day_results_ts.to_csv('data_summaries/audit_day_results_across_runs.csv')
fig, ax = plt.subplots(figsize=(22,3))
ax.plot(m_day_results_ts['bed_utilisation'])
ax.set_title('Bed Utilisation across model runtime (days)')
ax.set_ylabel('Mean daily proportion of bed utilisation')
return(fig)
11. Summary results per day for multiple runs#
Group by weekday
def weekly_summ_bed_utilisation(m_day_results):
"""
summarise per week across runs and save to csv in case of further analysis
print bed utilisation plot
warm-up results are excluded at runtime
"""
m_day_results_wd = m_day_results.groupby(['weekday']).mean()
m_day_results_wd.to_csv('data_summaries/audit_weekday_results_across_runs.csv')
values = m_day_results_wd['bed_utilisation']
names = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
fig, ax = plt.subplots(figsize=(8,4))
ax.bar(names, values)
ax.set_title('Mean bed Utilisation per day of week')
return(fig)
12. Patient level results summarised by day and weekday#
Lost slots calculation and plots per day and weekday
def calc_lost_theatre_slots(primary_pt_results, revision_pt_results):
"""
Join pt dataframes into single df
Select columns for lost slots
Summarise by day across runs
Deal with 0-day arrivals
Save to csv
Return lost_slots_df
"""
pt_results = pd.concat([primary_pt_results, revision_pt_results])
lost_slots_df = pt_results[["Day", "lost slots", "weekday"]]
lost_slots_df = pd.DataFrame(lost_slots_df.groupby(['Day', 'weekday'])['lost slots'].sum().astype(int))
lost_slots_df = lost_slots_df.assign(DayLostSlots = lambda x: (x['lost slots'] / DEFAULT_NUMBER_OF_RUNS))
lost_slots_df = pd.DataFrame(lost_slots_df["DayLostSlots"]).reset_index()
#0-arrival days excluded from df - add to Days sequence and fill lost slots value with 0 lost slots
# re-index as dataframe length increasing. Fill values in columns with 0.
lost_slots_df = (lost_slots_df.set_index('Day')
.reindex(range(lost_slots_df.Day.iat[0],lost_slots_df.Day.iat[-1]+1), fill_value=0)
.reset_index())
#change 0 weekdays into correct weekday integer
#need days of week seq and length of total range > length of dataframe
shortseq = np.arange(len(range(0,7)))
length = math.ceil(len(lost_slots_df) / 7)
#create total sequence and flatten array list into list of elements
sequence = ([np.tile((shortseq),length)])
flat_seq = list(itertools.chain(*sequence))
#truncate to correct length and save to column
sequence = flat_seq[:len(lost_slots_df)]
lost_slots_df['weekday'] = sequence
lost_slots_df.to_csv('data_summaries/Lost_slots_results_per_day.csv')
return(pd.DataFrame(lost_slots_df))
#lost_slots_df = calc_lost_theatre_slots(m_primary_pt_results, m_revision_pt_results)
def plot_lost_slots_per_day(lost_slots_df):
"""
Remove warm-up period results
Plot lost slots per day
"""
lost_slots_df = lost_slots_df[lost_slots_df["Day"] > DEFAULT_WARM_UP_PERIOD]
fig, ax = plt.subplots(figsize=(22,3))
ax.plot(lost_slots_df['DayLostSlots'])
ax.set_title('Lost theatre slots across model runtime (days)')
return(fig)
#plot_lost_slots_per_day(lost_slots_df);
def plot_lost_slots_per_week(lost_slots_df):
"""
Remove warm-up period results
Group by week
"""
lost_slots_df = lost_slots_df[lost_slots_df["Day"] > DEFAULT_WARM_UP_PERIOD]
lost_slots_wk_plot = lost_slots_df.groupby('weekday').mean()
lost_slots_wk_plot.reset_index()
values = lost_slots_wk_plot['DayLostSlots']
names = ['Monday', 'Tuesday', 'Wednesday', 'Thursday', 'Friday', 'Saturday', 'Sunday']
fig, ax = plt.subplots(figsize=(8,4))
ax.bar(names, values)
ax.set_title('Mean lost slots per day of week')
return(fig)
#plot_lost_slots_per_week(lost_slots_df);
13. MORE plots#
Determine appropriate replications
#calc means of multiple reps by rep
more_plot_results = m_day_results.groupby(['run']).mean()
more_plot_results_ = more_plot_results.loc[:, ['bed_utilisation','primary_bed_queue', 'revision_bed_queue',
'primary_mean_los', 'revision_mean_los']]
more_plot_results_ = more_plot_results_.reset_index(drop=True)
more_plot_results_ *= 100
more_plot_results_.columns = [0,1,2,3,4]
more_plot_results_.index.name = None
# more_plot_results_.head()
def ci_for_sample_mean(mean_value, std, n, critical_value=1.96):
'''Confidence interval for mean. Assume std is sample std.
Notes:
------
critical value hard coded at the moment.
Should update to use t dist.
'''
half_width = (critical_value * (std / np.sqrt(n)))
mean_lower = mean_value - half_width
mean_upper = mean_value + half_width
return mean_lower, mean_upper
def ci_percentile(results, field, percentile, critical_value=1.96):
'''Approximate confidence interval for percentile.
Note these may or may not be symmetric.
Notes:
------
critical value hard coded at the moment.
Should update to use t dist.
Params:
------
results: pd.DataFrame
Results dataframe - tabular data where each row is a rep and each col is a KPI
field: int
Field from data frame to analyse
percentile: float
The percentile around which to form the CI
critical_value: float, optional (default = 1.96)
critical value of the normal dist to use.
'''
half_width = critical_value * np.sqrt((percentile * (1 - percentile)) / (len(results) - 1))
y_beta_1 = results[field].quantile(percentile - half_width)
y_beta_2 = results[field].quantile(percentile + half_width)
return y_beta_1, y_beta_2
def as_horizontal_axis_fraction(value, xmin, xmax):
'''Convert a axis value to a fraction accounting for the
minimum on the xaxis (i.e. axis may not start from 0).
'''
return (value - xmin) / (xmax - xmin)
def draw_interval(ax, start, end, style="|-|", lw=3, color='b'):
'''Annotate a matplotlib chart underneath x axis with an confidence interval.
'''
_ = ax.annotate('', xy=(start, -0.1), xycoords='axes fraction',
xytext=(end, -0.1),
arrowprops=dict(arrowstyle=style, color=color, lw=lw))
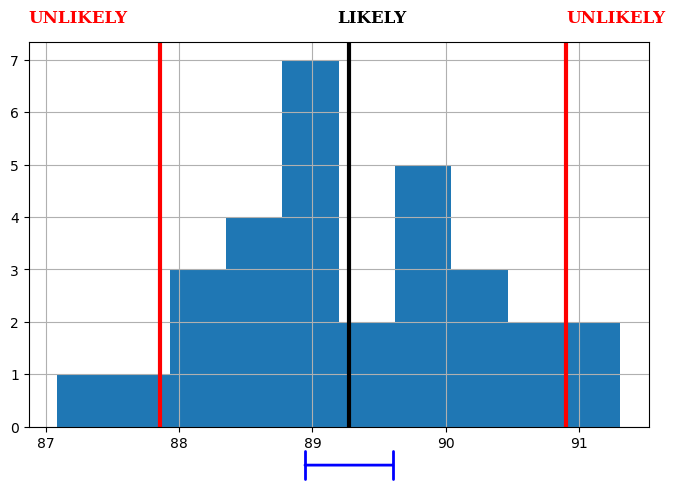
def more_plot(results, field=0, bins=None, figsize=(8, 5), percentiles=(0.05, 0.95), surpress_warnings=False):
'''Measure of Risk and Error (MORE) plot.
Risk illustrated via likely and unlikely ranges of replication values.
Erorr illustrated for CIs for mean and wide approx confidence intervals for percentiles
Confidence intervals for percentiles will only be calculated if > 80 replications due to
approximation accuracy.
Notes:
------
Each value plotted represents the mean of a replication (e.g. daily throughput). It should
not be confused with an individuals results (e.g. an individuals throughput time).
If the system modelled contains time dependency the MORE plot may hide time of day/event effects.
Params:
------
results: pd.DataFrame
Tabular data of replications. each column is a kpi
field: int
ID of column containing relevant data
bins: int, optional (default=None)
no. bins to generate. None=pandas decides no.
figsize: tuple, optional (default=(8,5))
size of plot
Returns:
-------
fig, ax
Refs:
-----
Nelson 2008. (Winter Simulation Paper)
https://ieeexplore.ieee.org/document/4736095
'''
# probably will shift these to module level scope.
LIKELY = 'LIKELY'
UNLIKELY = 'UNLIKELY'
FONT_SIZE = 12
LINE_WIDTH = 3
LINE_STYLE = '-'
CRIT_VALUE = 1.96
UPPER_QUANTILE = percentiles[1]
LOWER_QUANTILE = percentiles[0]
INTERVAL_LW = 2
MIN_N_FOR_PERCENTILE = 80
WARN = f'CIs for percentiles are not generated as sample size < {MIN_N_FOR_PERCENTILE}.'
WARN += ' To supress this msg set `supress_warnings=True`'
ax = results[field].hist(bins=bins, figsize=figsize)
mean = results[field].mean()
std = results[field].std(ddof=1)
upper_percentile = results[field].quantile(UPPER_QUANTILE)
lower_percentile = results[field].quantile(LOWER_QUANTILE)
# vertical lines
ax.axvline(x=mean, linestyle='-', color='black', linewidth=LINE_WIDTH)
ax.axvline(x=upper_percentile, linestyle='-', color='red', linewidth=LINE_WIDTH)
ax.axvline(x=lower_percentile, linestyle='-', color='red', linewidth=LINE_WIDTH)
like_font = {'family': 'serif',
'color': 'black',
'weight': 'bold',
'size': FONT_SIZE
}
unlike_font = {'family': 'serif',
'color': 'red',
'weight': 'bold',
'size': FONT_SIZE
}
# add text
txt_offset = ax.get_ylim()[1] * 1.05
ax.text(mean - (mean * 0.001), txt_offset, LIKELY, fontdict=like_font)
ax.text(upper_percentile, txt_offset, UNLIKELY, fontdict=unlike_font)
ax.text(ax.get_xlim()[0], txt_offset, UNLIKELY, fontdict=unlike_font)
# calculate and display confidence intervals
## CIs for sample mean
mean_lower, mean_upper = ci_for_sample_mean(mean, std, len(results))
# Draw Confidence intervals
# The horizontal lines are expressed as an axis fraction i.e. between 0 and 1.
# This means thatthe percentile CIs need to be converted before plotting.
# The function as_horizontal_axis_fraction is used.
## mean CI
hline_mean_from = as_horizontal_axis_fraction(mean_lower, ax.get_xlim()[0], ax.get_xlim()[1])
hline_mean_to = as_horizontal_axis_fraction(mean_upper, ax.get_xlim()[0], ax.get_xlim()[1])
draw_interval(ax, hline_mean_from, hline_mean_to, lw=INTERVAL_LW)
# avoid approximation issues with small samples.
if len(results) >= MIN_N_FOR_PERCENTILE:
## upper percentile
y_beta_1, y_beta_2 = ci_percentile(results, field, UPPER_QUANTILE, critical_value=CRIT_VALUE)
## lower percentile
y_beta_l_1, y_beta_l_2 = ci_percentile(results, field, LOWER_QUANTILE, critical_value=CRIT_VALUE)
## line for upper quantile CI
hline_upper_q_from = (y_beta_1 - ax.get_xlim()[0]) / (ax.get_xlim()[1] - ax.get_xlim()[0])
hline_upper_q_to = (y_beta_2 - ax.get_xlim()[0]) / (ax.get_xlim()[1] - ax.get_xlim()[0])
hline_upper_q_from = as_horizontal_axis_fraction(y_beta_1, ax.get_xlim()[0], ax.get_xlim()[1])
hline_upper_q_to = as_horizontal_axis_fraction(y_beta_2, ax.get_xlim()[0], ax.get_xlim()[1])
draw_interval(ax, hline_upper_q_from, hline_upper_q_to, lw=INTERVAL_LW)
## line for lower quantile CI
hline_lower_q_from = (y_beta_l_1 - ax.get_xlim()[0]) / (ax.get_xlim()[1] - ax.get_xlim()[0])
hline_lower_q_to = (y_beta_l_2 - ax.get_xlim()[0]) / (ax.get_xlim()[1] - ax.get_xlim()[0])
hline_lower_q_from = as_horizontal_axis_fraction(y_beta_l_1, ax.get_xlim()[0], ax.get_xlim()[1])
hline_lower_q_to = as_horizontal_axis_fraction(y_beta_l_2, ax.get_xlim()[0], ax.get_xlim()[1])
draw_interval(ax, hline_lower_q_from, hline_lower_q_to, lw=INTERVAL_LW)
elif not surpress_warnings:
warnings.warn(WARN)
return ax.figure, ax
results = more_plot_results_
fig, ax = more_plot(results);
/tmp/ipykernel_254019/625959835.py:126: UserWarning: CIs for percentiles are not generated as sample size < 80. To supress this msg set `supress_warnings=True`
warnings.warn(WARN)
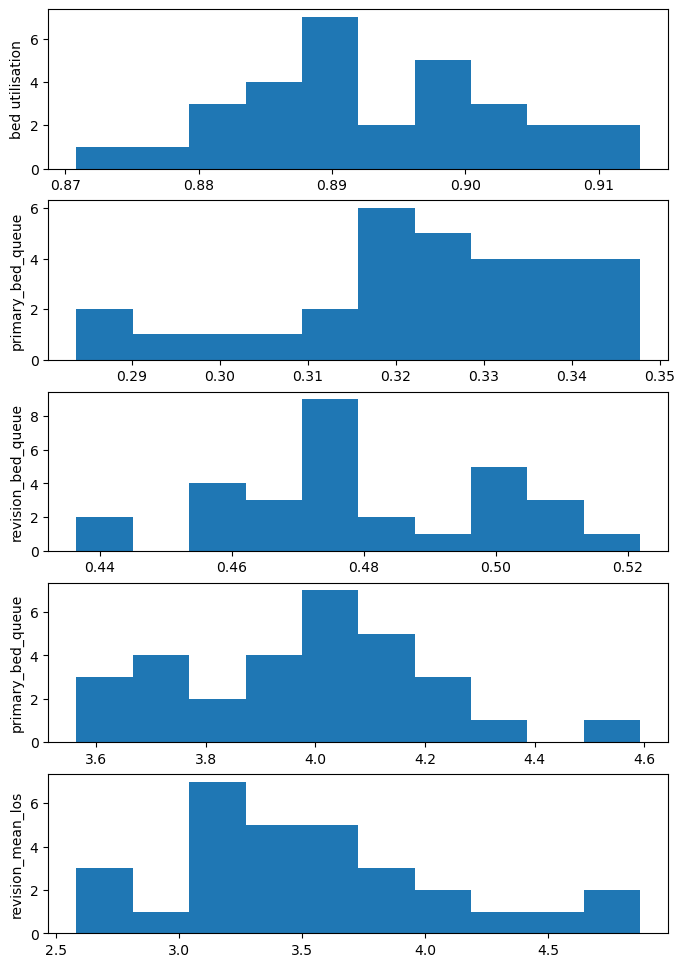
hists of outputs#
more_plot_results = m_day_results.groupby(['run']).mean()
hist_results = more_plot_results.loc[:, ['bed_utilisation', 'primary_bed_queue', 'revision_bed_queue',
'primary_mean_los', 'revision_mean_los']]
fig, ax = plt.subplots(5, 1, figsize=(8,12))
ax[0].hist(hist_results['bed_utilisation']);
ax[0].set_ylabel('bed utilisation')
ax[1].hist(hist_results['primary_bed_queue']);
ax[1].set_ylabel('primary_bed_queue');
ax[2].hist(hist_results['revision_bed_queue']);
ax[2].set_ylabel('revision_bed_queue');
ax[3].hist(hist_results['primary_mean_los']);
ax[3].set_ylabel('primary_bed_queue');
ax[4].hist(hist_results['revision_mean_los']);
ax[4].set_ylabel('revision_mean_los');